 Evidence for sequential and increasing activation of replication origins along replication timing gradients in the human genome. Open chromatin encoded in DNA sequence is the signature of ‘master’ replication origins in human cells. Replication-associated mutational asymmetry in the human genome. Human gene organization driven by the coordination of replication and transcription. Replication-associated strand asymmetries in mammalian genomes: toward detection of replication origins. Asymmetric substitution patterns in the two DNA strands of bacteria. Genetic variation in human DNA replication timing. High-resolution Repli-Seq defines the temporal choreography of initiation, elongation and termination of replication in mammalian cells. Impact of replication timing on non-CpG and CpG substitution rates in mammalian genomes. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Bubble-seq analysis of the human genome reveals distinct chromatin-mediated mechanisms for regulating early- and late-firing origins. Bubble-chip analysis of human origin distributions demonstrates on a genomic scale significant clustering into zones and significant association with transcription. Genome-wide identification and characterisation of human DNA replication origins by initiation site sequencing (ini-seq. Allele-specific genome-wide profiling in human primary erythroblasts reveal replication program organization. Genomic study of replication initiation in human chromosomes reveals the influence of transcription regulation and chromatin structure on origin selection. Unraveling cell type-specific and reprogrammable human replication origin signatures associated with G-quadruplex consensus motifs. Genome-wide studies highlight indirect links between human replication origins and gene regulation. Dynamics of DNA replication in mammalian somatic cells: nucleotide pool modulates origin choice and interorigin spacing. Regulation of DNA replication within the immunoglobulin heavy-chain locus during B cell commitment. DNA replication origin interference increases the spacing between initiation events in human cells. Lebofsky, R., Heilig, R., Sonnleitner, M., Weissenbach, J. Genomic methods for measuring DNA replication dynamics. Peaks cloaked in the mist: the landscape of mammalian replication origins. On the mechanism of DNA replication in mammalian chromosomes. Besides revealing the genome replication program in fine detail, OK-seq has been instrumental in numerous studies unravelling mechanisms of genome stability, epigenome maintenance and genome evolution. The experiments and the data processing can be accomplished within six days. These tools are essential for the accurate interpretation of human and yeast replication programs. Furthermore, we present the analytical approach based on a hidden Markov model, which allows automated detection of ascending, descending and flat replication fork directionality segments revealing the zones of replication initiation, termination and unidirectional fork movement across the entire genome. Here we provide the detailed experimental procedures for performing OK-seq in unperturbed cultured human cells and budding yeast and the bioinformatics pipelines for data processing and computation of replication fork directionality. OK-seq quantitates the proportion of leftward- and rightward-oriented forks at every genomic locus and reveals the location and efficiency of replication initiation and termination events. 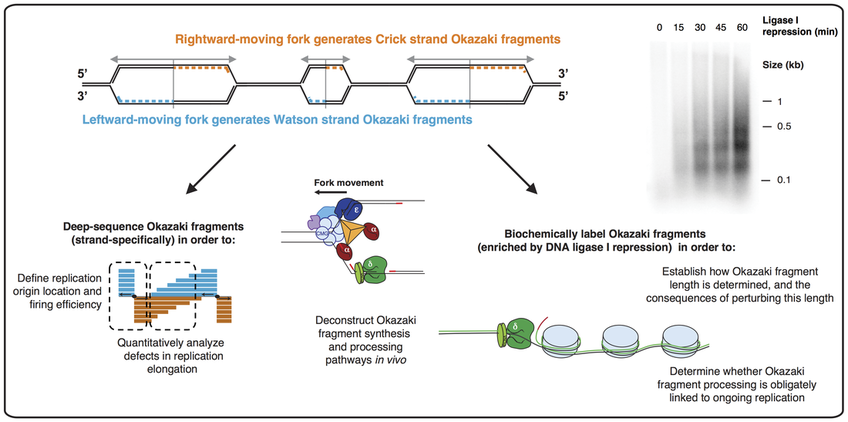  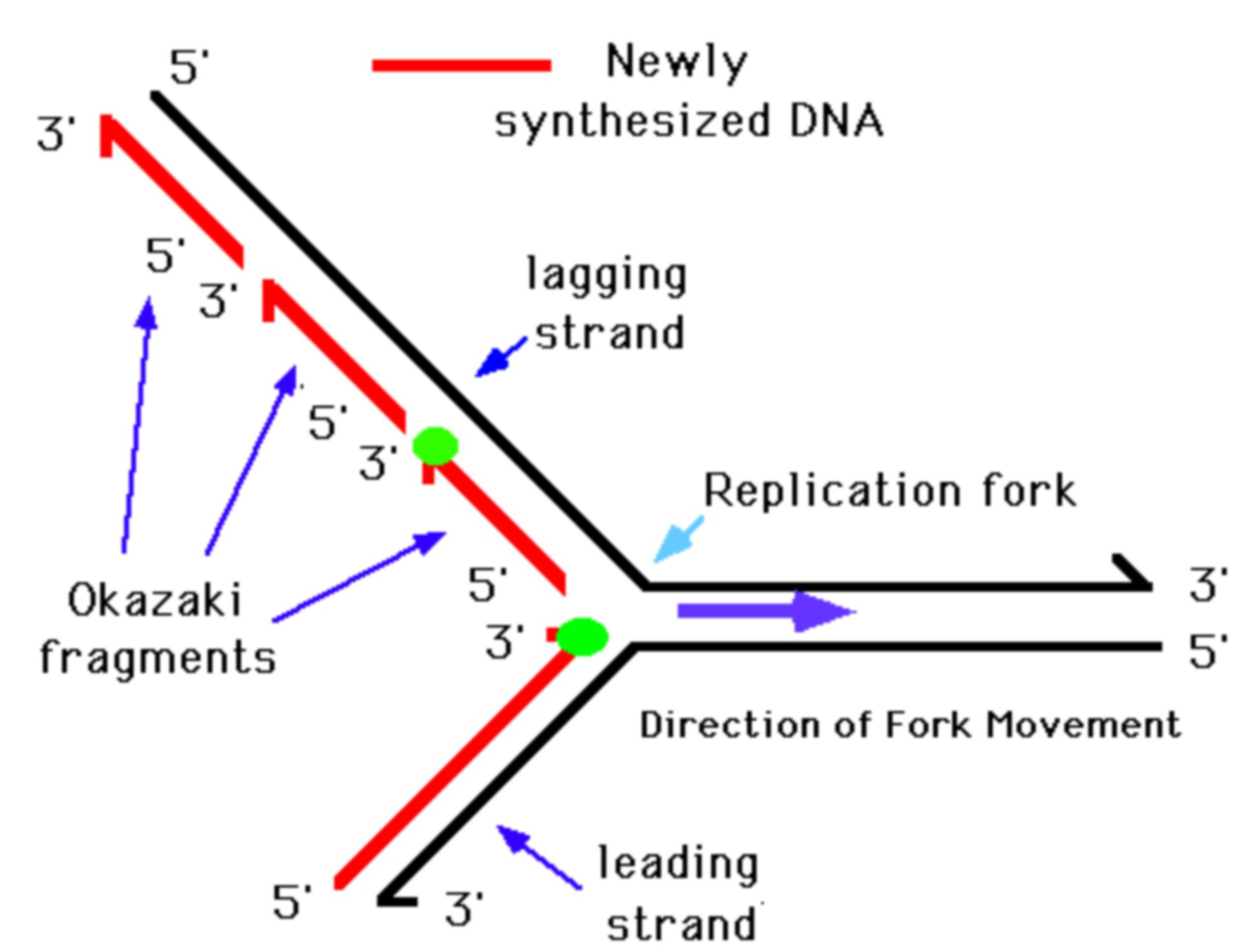
To reveal the location of replication initiation and termination in the human genome, we developed Okazaki fragment sequencing (OK-seq), a quantitative approach based on the isolation and strand-specific sequencing of Okazaki fragments, the lagging strand replication intermediates. Studying the dynamics of genome replication in mammalian cells has been historically challenging. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
